Biome-BGC
Project Summary
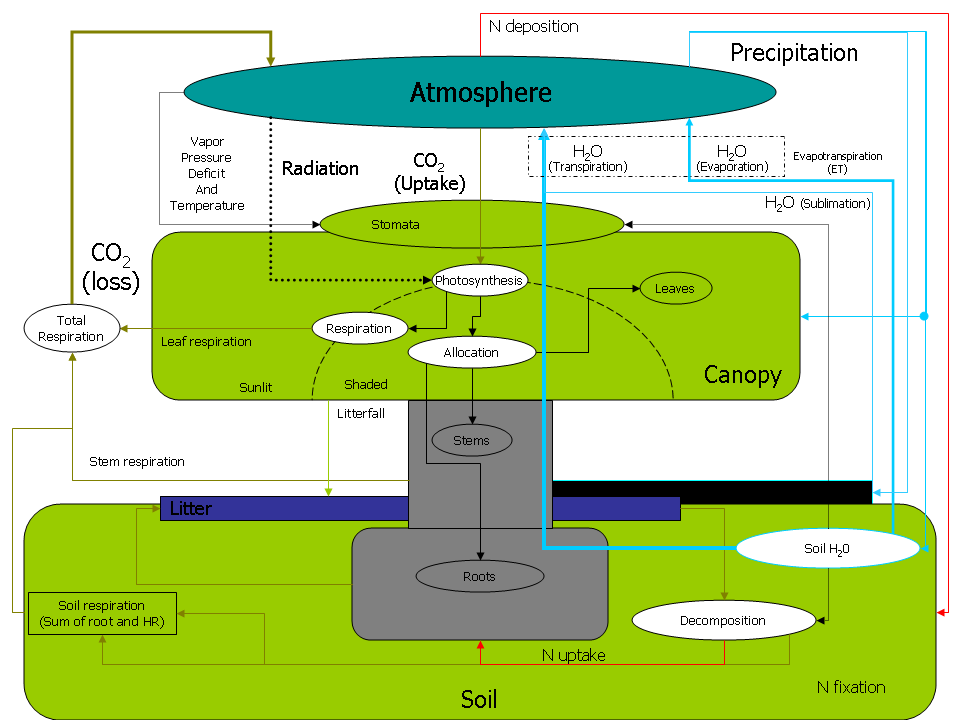
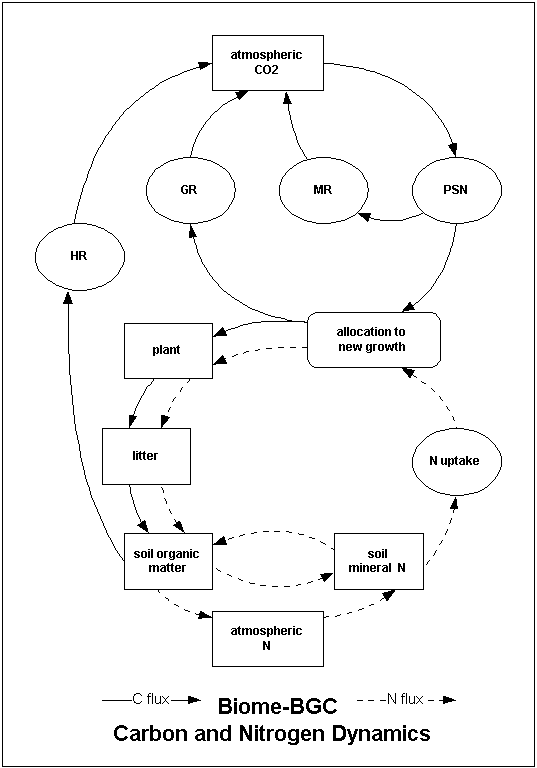
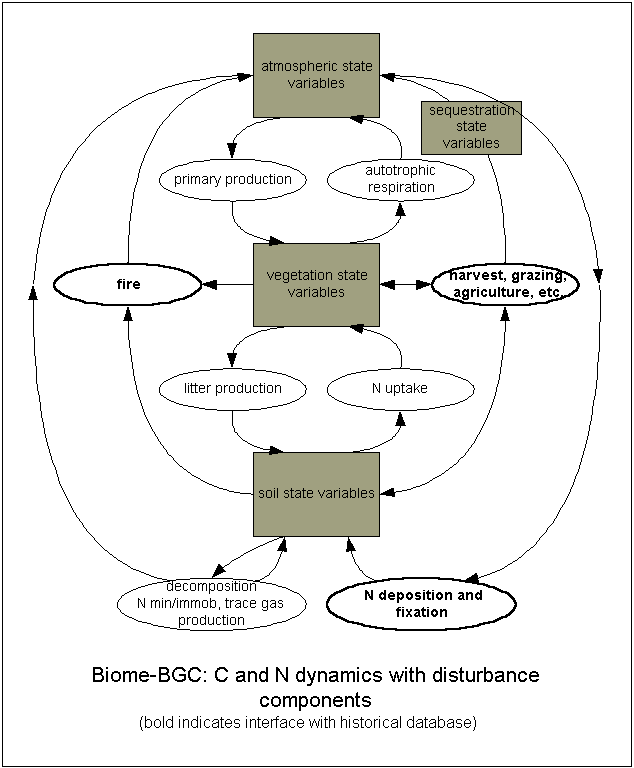
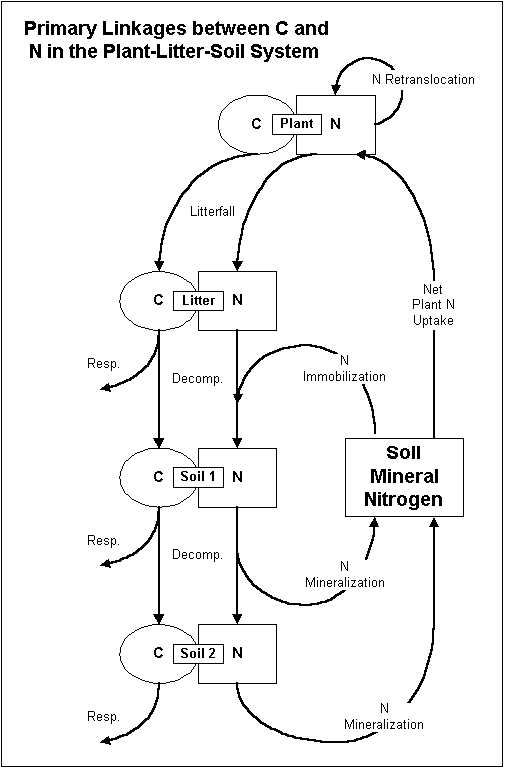
 This project page serves as a central place for model information and source code. Biome-BGC is an ecosystem process model that estimates storage and flux of carbon, nitrogen and water. Biome-BGC is a computer program that estimates fluxes and storage of energy, water, carbon, and nitrogen for the vegetation and soil components of terrestrial ecosystems.
This project page serves as a central place for model information and source code. Biome-BGC is an ecosystem process model that estimates storage and flux of carbon, nitrogen and water. Biome-BGC is a computer program that estimates fluxes and storage of energy, water, carbon, and nitrogen for the vegetation and soil components of terrestrial ecosystems.
We call it a process model because its algorithms represent physical and biological processes that control fluxes of energy and mass. These processes include:
- New leaf growth and old leaf litterfall
- Sunlight interception by leaves, and penetration to the ground
- Precipitation routing to leaves and soil
- Snow accumulation and melting
- Drainage and runoff of soil water
- Evaporation of water from soil and wet leaves
- Transpiration of soil water through leaf stomata
- Photosynthetic fixation of carbon from CO2 in the air
- Uptake of nitrogen from the soil
- Distribution of carbon and nitrogen to growing plant parts
- Decomposition of fresh plant litter and old soil organic matter
- Plant mortality
- Fire
The model uses a daily time-step. This means that each flux is estimated for a one-day period. Between days, the program updates its memory of the mass stored in different components of the vegetation, litter, and soil.
Weather is the most important control on vegetation processes. Flux estimates in Biome-BGC depend strongly on daily weather conditions. Model behavior over time depends on the history of these weather conditions, the climate.
Biome-BGC: Required meteorological inputs
The primary driving variables for estimating ecosystem processes with Biome-BGC are daily meteorological data. One of the main premises of the Biome-BGC logic is that if we can adequately describe the dependence of key processes on environmental factors related to daily meteorology, then we can estimate these processes anywhere that daily meteorological data are available.
The following near-surface meteorological parameters are required by Biome-BGC:
- Daily maximum temperature (°C)
- Daily minimum temperature (°C)
- Daylight average temperature (°C)
- Daily total precipitation (cm)
- Daylight average partial pressure of water vapor (Pa)
- Daylight average shortwave radiant flux density (W/m2)
- Daylength (s)
In many cases, the only data available for a particular site are the daily maximum and minimum temperatures, and the daily total precipitation. We have developed simple and accurate algorithms to estimate the radiation and humidity parameters when they are missing (see our MT-CLIM page).
In other cases, we are interested in performing simulations at a site that has no meteorological measurements at all. In this case, if there is a nearby weather station, perhaps at a different elevation, then MT-CLIM is used to estimate the site meteorology. Often there is not a station very nearby, or there are two or more neighboring stations that might be used to estimate the daily meteorological parameters for the site. In that case it may be necessary to interpolate between individual stations, and possibley extrapolate to a different elevation. This is often the case when we perform gridded Biome-BGC simulations over large regions. We have developed a sophisticated set of algorithms to treat this problem (see our Daymet page).
Code & Documentation
Get the latest code from GitHub
Download Biome-BGC 4.2 from NTSG
Biome-BGC User Guide | Biome-BGC Theoretical Basis
Biome-BGC version 5.0 Release>
Download the latest Biome-BGC (Biome-BGCMUSO v5.0) model code and documentation
Biome-BGC version 4.2 Final Release>
We have been hard at work compiling a list of bug fixes and model enhancments into a new version of Biome-BGC. We are offereing this new beta model version under the downloads section of this site. We have been hard at working compiling a list of bug fixes and model enhancments into a new version of Biome-BGC. We are offereing this new beta model version under the downloads section of this site.
This new version has a number of minor bug fixes and some increased functionality. The following is a list of relavant changes to this version.
Development Notes for Biome BGC. This file is a list of code/internal changes.
4.2rc2
- Fixed Spin & Go so that the '-g' flag actually works.
- IMPORTANT Corrected use of the co2 file. It now correctly looks up the value for the model year. Previously it simply used co2 entries starting at the top of the co2 file.
- Added External Nitrogen Deposition File see USAGE.TXT for more information.
4.2rc1
- Identical to last 4.2pre
4.2pre
a. Added command line option support using getopts() to add functionality without breaking backwards compatibility. See USAGE.TXT for options.
b. Centralized header files bgc.h for libbgc pointbgc.h for pointbgc C source files now only need to include one header, all others get included automatically in the correct order.
c. Added bgc_io.c for non-data I/O. Replaced all printf()s to stdout with the new wrapper function bgc_printf().
a. bgc_printf() is for outputing information to standard out and errror it behaves similar to printf but with an extra first argument:
-
- BV_ERROR: For fatal error messages
- BV_WARN: For non-fatal warnings
- BV_PROGRESS: For basic progress information
- BV_DETAIL: For detailed progress information
- BV_DIAG: For internal diagnostic/debug messages
- VERBOSITY is one of several defined keywords. You use the keyword appropriate to when your message should be printed. bgc_printf also supports logging functionality see USAGE.TXT for usage.
- Replaced all DEBUG printf's with proper bgc_printf calls
- Merged Spinup code into main code to reduce code duplication merged spinup_bgc.c into bgc.c
- Added pan arctic summary output. Traditional behavior is set by signed char summary_sanity = INSANE ; in bgc()
- SANE behavior (pan-artic style) calculates these correclty: Maximum Monthly LAI End of month Snow Water End of month Sail Water content
- Also added new output_map variables
- output_map[641] = &summary->daily_et;
- output_map[642] = &summary->daily_outflow;
- output_map[643] = &summary->daily_evap;
- output_map[644] = &summary->daily_trans
- output_map[645] = &summary->daily_soilw;
- output_map[646] = &summary->daily_snoww;
- Rewrote make files to not use version specific directories
- Added 'make test' to build then run a spinup and model test
- Moved all photosynthesis science code out of bgc.c and into photosynthesis.c
- Corrected some sprintf formating to be more ANSI C standards complient in prephenology.c
- Added tests/corrections for negative variables in radtrans.c and metarr_init.c: (swavgfd,par,swabs_plaishade,parabs_plaishadei,dal)
- Added code to improve portability and standards compliance.
- Added 'make tools' to master makefile. This builds the restart diff/view tool.
- Replaced floating point equality tests with tolerance tests. Global tolerance define: FLT_COND_TOL = 1e-10 epc_init.c, sitec_init.c
- Various memory leak and pointer assignment fixes in pointbgc.c
- Fix to prevent SEGFAULT on empty restart file in pointbgc.c
- Split output_init.c into output_ctrl.c and output_init.c output_ctrl.c reads the output controls from the ini file output_init.c initializes the output files
- Added 'spin-n-go' functionality. This mode '-g' flag will run both the model and the spinup without generating the restart file.
NTSG Personnel
- Steve Running
- Jordan Golinkoff
- Ryan Anderson